How to import ETER data into R
The R programming language is well suited to work with the ETER database. We, the ETER project team, for example use R for data validation, data quality and also for creating the analytical reports. In this blog post, we will guide you on importing the ETER data into R. We assume, that you downloaded the whole dataset as .csv-file (recommended) and that you have installed R already. We use the open-source IDE RStudio for scripting and executing R functions.
Please open R, RStudio or any other editor you may use. We will import the ETER data withouth additional R packages and only need the R base package, which means that we don’t have to load any additional libraries.
Beforehand you need to know that value fields within the ETER data can contain value codes. This means that a value in ETER can either be a number or a code which give you some additional information (e.g. “a” means not applicable, “m” means missing etc.; see a full list of all value codes here). Thus, we have the possibility of importing the dataset in two different ways:
- Importing the data as they are, showing either numbers or value codes. Mixed columns are of class “character” in R and cannot be used for calculations.
- Importing value codes as NA (not available). This has the advantage that the concerned columns are of type “numeric” and can be used for calculations
How to import ETER data into R
The R programming language is well suited to work with the ETER database. We, the ETER project team, for example use R for data validation, data quality and also for creating the analytical reports. In this blog post, we will guide you on importing the ETER data into R. We assume, that you downloaded the whole dataset as .csv-file (recommended) and that you have installed R already. We use the open-source IDE RStudio for scripting and executing R functions. Please open R, RStudio or any other editor you may use. We will import the ETER data withouth additional R packages and only need the R base package, which means that we don’t have to load any additional libraries. Beforehand you need to know that value fields within the ETER data can contain value codes. This means that a value in ETER can either be a number or a code which give you some additional information (e.g. “a” means not applicable, “m” means missing etc.; see a full list of all value codes here). Thus, we have the possibility of importing the dataset in two different ways:
- Importing the data as they are, showing either numbers or value codes. Mixed columns are of class “character” in R and cannot be used for calculations.
- Importing value codes as NA (not available). This has the advantage that the concerned columns are of type “numeric” and can be used for calculations.
First, we are importing the data in the format they are displayed in the database. This enables the user to get all the information coming from special codes and is also the format we use for data validation, where we check the data with respect to their internal consistency. In order to work with the dataset, we need to assign a name (data), when we import using the read.csv2-function: data <- read.csv2(“YOURDATAPATH/eter_export_all.csv”, sep = “;”, dec = “,”, header = TRUE, na.strings = “”, quote = “””, comment.char = “”, colClasses = “character”, encoding = “UTF-8”) Since the header of the first column (ETER ID + Year) is not named correctly when importing the file in R, we need to update the column name separately: names(data)[1] <- “ETERID.Year” When you have a detailed look at the data now, you will see that the data are displayed in R as they are in the database: data[500:510, c(2, 5, 6, 361:363)] 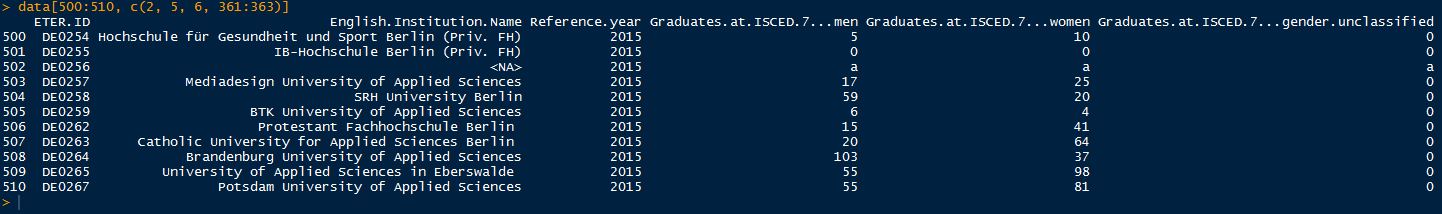 Congratulations, you have now imported the ETER data into R. This is the choice if you want to have the value codes in your dataset, e.g. to do data validation checks or just simply have a detailed look at them. If you want to do also calculations, then you need to get rid of the value codes in order to enable R to treat the number columns as numeric. In order to do so, you can import the data in the following way (we name the dataset data_numeric in this case): data_numeric <- read.csv2(“YOURDATAPATH/eter_export_all.csv”, sep = “;”, dec = “,”, header = TRUE, stringsAsFactors = FALSE, na.strings = c(“”, “a”, “m”, “x”, “xr”, “xc”,”nc”, “x “, “m “), quote=”””, comment.char = “”,encoding = “UTF-8”) The key in this function is to define the value codes (e.g. “a”, “m”, “x”, etc.) and the empty cells (“”) as NAs, which enables R to treat the columns as numeric (remember, R cannot define a column of class “numeric”, if characters are in there, so we have to get rid of them). Again, we need to rename the first column: names(data_numeric)[1] <- “ETERID.Year” You can now compare the outcome of the import command to the former import above. Value codes are now turned to NA instead of “a”. data_numeric[500:510, c(2, 5, 6, 361:363)]
Congratulations, you have now imported the ETER data into R. This is the choice if you want to have the value codes in your dataset, e.g. to do data validation checks or just simply have a detailed look at them. If you want to do also calculations, then you need to get rid of the value codes in order to enable R to treat the number columns as numeric. In order to do so, you can import the data in the following way (we name the dataset data_numeric in this case): data_numeric <- read.csv2(“YOURDATAPATH/eter_export_all.csv”, sep = “;”, dec = “,”, header = TRUE, stringsAsFactors = FALSE, na.strings = c(“”, “a”, “m”, “x”, “xr”, “xc”,”nc”, “x “, “m “), quote=”””, comment.char = “”,encoding = “UTF-8”) The key in this function is to define the value codes (e.g. “a”, “m”, “x”, etc.) and the empty cells (“”) as NAs, which enables R to treat the columns as numeric (remember, R cannot define a column of class “numeric”, if characters are in there, so we have to get rid of them). Again, we need to rename the first column: names(data_numeric)[1] <- “ETERID.Year” You can now compare the outcome of the import command to the former import above. Value codes are now turned to NA instead of “a”. data_numeric[500:510, c(2, 5, 6, 361:363)] 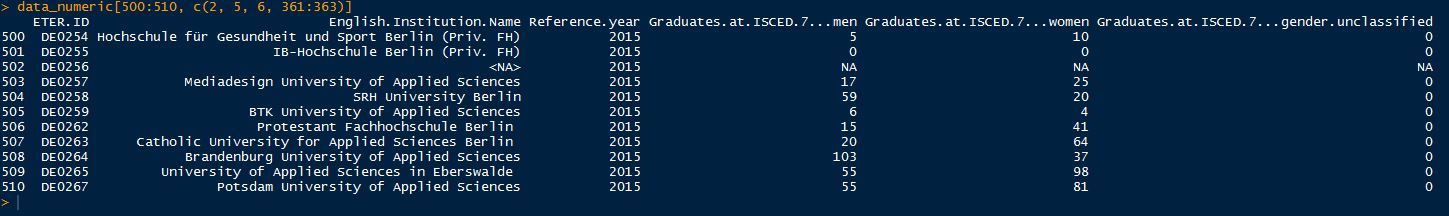
You are now ready to use the full variety of the ETER dataset for your research. If you have questions to this post, please contact us at eter@eter-project.com. Also, if you have additional questions on the ETER project, technical or not, do not hesitate to contact us.
